 If you have more than 5 or 6 sequences to model, it is easier for you. I think such a resource would be more of a hindrance than an asset to the scientific community. as well as models downloaded from the AlphaFold Protein Structure Database Info icon. If there was a database of predicted secondary structures, people would likely take them for granted (make the equation prediction = fact) which would be quite "unscientific". Also they change each time the algorithms are improved. 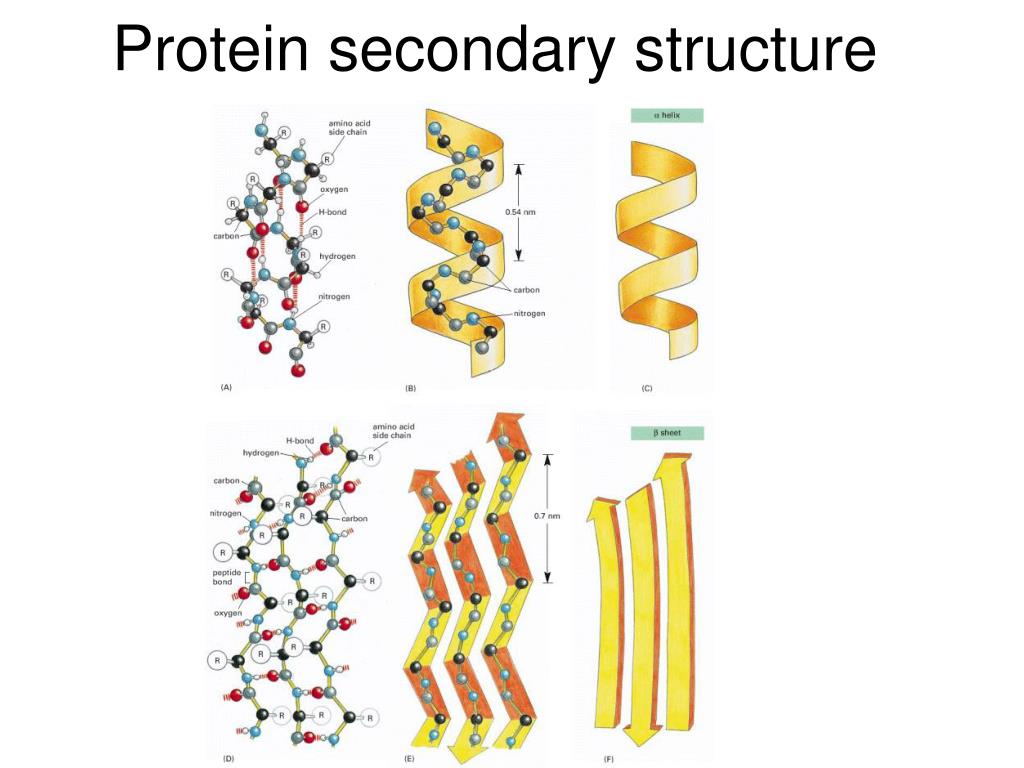
The results may differ from one prediction method to another. 1.PRIMARY STRUCTURE: Primary proteins are unbranched polymers of amino acids linked head. They're "educated guesses" that are often correct, but sometimes wrong. Given a query amino acid sequence, CRNPRED predicts one-dimensional structures including secondary structures, contact numbers and residue-wise contact orders. Protein secondary structures have been identified as the links in the physical processes of primary sequences, typically random coils, folding into functional tertiary structures that enable proteins to involve a variety of biological events in life science. And I use them quite frequently: they are extremely useful! But as far as I know, no prediction is accepted as fact. In those cases, it is not only the secondary structures but also the tertiary structures (with the caveat that the crystal structure of a protein does not prove "all" states that a protein may take in real "dynamic" physiological conditions).įor all those proteins that have not been crystalized, then we can only rely on predictions. The two common secondary structures encountered in proteins are alpha-helix ( -helix and -pleated sheet. 
That may be the case for secondary structures of proteins, but only in the case where the said proteins have been crystalized. Protein structure is often described at four different scales. The second level of protein structure, secondary structure, is the local conformation adopted by stretches of contiguous amino acids. I'm not sure that makes much sense, let me explain my point of view (which may not be that of everyone): most often, the data that is found in databases is the "state of knowledge" of the described object, based on experimentation. You suggest that there should be a Secondary Structure Database. 
I think you found the best answer yourself: use a predictor! There are several out there. In addition to protein secondary structure, JPred also makes predictions of solvent accessibility and coiled-coil regions. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
